SPANA for Spectral Data Analyses [Analyses] Copyright (c) 2010 ykuroda All right reserved. |
1. Equilibrium and Reaction Rate Analyses
SPANA performs the equilibrium or rate analyses for the
titration data or time course data by using the non-linear least square
optimization procedure.
The calculation minimizes the residual square sum value (ss) to optimize
the parameters such as the equilibrium constants, rate constants and absorptivities.
The ss value is evaluated from the observed strengths of spectra (Yobs) and the theoretical ones (Ytheoretical) calculated by using the parameters and independent experimental variable
such as concentrations, temperature and/or time, as follows.
ss = ∑(Yobs - Ytheoretical)^2
SPANA provides the following equilibrium and rate models.
| Model | Theoretical Spectral Strength |
| Equilibrium Model | |
| Sequential 1:n Complex formationl ABi-1 + B <-> ABi , Ki= [ABi]/[ABi-1][B] i = 1~n , nmax = 5 |
Abs = εA[A] + ∑εABi[ABi] |
| Sequential 1:n Complex formation accompaning Pre-equilibrium of A pA* <-> qA, Ko= [A]q / [A*]p ABi-1 + B <-> ABi , Ki= [ABi]/[ABi-1][B] i = 1~n , nmax = 5 |
Abs = εA*[A*] + εA[A] + ∑εABi[ABi] |
| Independent 1:n Complex formation Aempty i Bj-1 + B <-> Aoccupied i Bj Ki = [Aoccupied i Bj]/[Aempty i Bj-1][B] i , j = 1~n , nmax = 5 |
Abs = (∑εA)[A] + ∑( ∑emptyεA + ∑occupiedεAiB)[ABj] |
| pH titration model with n pKa’s AHi-1 + H+ <-> AHi Ki= [AHi]/[AHi-1][H+] i = 1~n , nmax = 5 |
Abs = εA[A] + ∑εAHi [AHi] |
| n-Step 1 : n Complex formation ABi-1 + B <-> ABi K=[ABi]/[ABi-1][B]a (i = 1~n) (a=1 : Independent complexation (a>1 : Cooperative complexation <Hill’s type expression>) - Job's Analyses - [A]Total + [B]Total = C (constant) φ = [B]Total / C , 1 - φ = [A]Total / C , φ= 0 ~1 |
Abs = nεA[A] + ∑{(n - i)εA + iεAB}(n ! / ((n - i) !・i !)) [ABi] |
| One-step m:n Complex formation mA + nB <-> AmBn K = [AmBn]/[A]m[B]n - Job's Analyses - [A]Total + [B]Total = C (constant) φ = [B]Total / C , 1 - φ = [A]Total / C , φ= 0 ~1 |
Abs = εA[A] + εAmBn[AmBn] |
| One-step m-n Self-reorganizaton mA <-> nA* K = [A*]n/[A]m |
Abs = εA[A] + εA*[A*] |
| Thermal Dependency mA <-> nA* K = [A*]n/[A]m = exp(-(ΔH-TΔS)/RT) A + B <-> AB K = [AB]/[A][B] = exp(-(ΔH-TΔS)/RT) |
Abs = εA[A] + εA*[A*] Abs = εA[A] + εAB[AB] |
| Competitive 1:1 Complex formation A + I <-> AI K0 = [AI]/[A][I] A + B <-> AB K1 = [AB]/[A][B] A, B : spectroscopically inactive I : spectroscopically active |
Abs = εI [I] + εAI [AI] |
| Competitive 1:2 Complex formation A + I <-> AI K0 = [AI]/[A][I] A + B <-> AB K1 = [AB]/[A][B] AB + B <-> BAB K2 = [BAB]/[AB][B] A, B : spectroscopically inactive I : spectroscopically active |
Abs = εI [I] + εAI [AI] |
| Reaction Rate Model | |
| 0 th Oder reaction |
Abs = k t +Abs0 |
| 1st Order reaction A → B k : rate const. |
Abs = εA[A]t + εB[B]t |
| 2nd Order Reaction A + B → C k : rate const. |
Abs = εA[A]t + εB[B]t + εC[C]t |
| Successive Reaction A → B → C k1, k2 : rate const. |
Abs = εA[A]t + εB[B]t + εC[C]t |
| NMR Line Shape Analyses A <-> B k+ , k- : rate const. |
I(v) = f (v, vA, vB, pA, k+, T2A, T2B, C) |
| General Function y : dependent variable t : independent variable a, b, c, d, e, f : parameters to be optimized |
y = a √ t + b y = a t + b y = a t2 + b t + c y = a t3 + b t2 + c t + d y = a t4 + b t3 + c t2 + d t + e y = a t5 + b t4 + c t3 + d t2 + e t + f y = (a t + b) / (t + c) + dt y = a ln (b t) + c t + d y = a exp (b t) + c t + d y = a exp (b t) + c exp (d t) + e y = a / (1 + exp (-(t - b) /c) + d t + e (Logistic function) y = a / (1 + (t - b)2/c2) + d t + e (Lorentz function) y = a exp (-(t - b)2/c2) +d t + e (Gauss function) |
| User's Function y[i] : dependent variable x[1] : independent variable ( = t) c[i] : constant defined in the program p[i] : parameters to be optimized |
y[i] = f ( x[1], c[i], p[i] ) |
2. Wave Separation
SPANA perform wave separation using Gauss, Lorentz, and Voigt functions.
The type and number of the wave components are set by the user.
3. Component analyses
SPANA estimates the contents of the standard spectra of components (max.
10) in the observed spectra by the method of multiple regression analysis.
4. Least Square Analyses for User's Functions
The least square analyses for user defining functions are available. The
following are the outline of preparation of the program which can be used
in SPANA (for the practical procedures of the analyses, see "Example"
page).
a) The Data
SPANA produces the text file, spana_lsq..dta, containing the collected
spectral data in the spana folder. The file format is shown below.
Format of "spana_lsq.dta" file
sp : number of sampling point, ds : number of data set, d(i) [ i = 1
- ds ] : number of data for ith data set
y(j) : spectral strength , t : condition value,
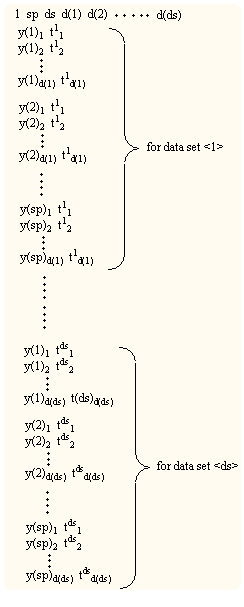
b) The Program
SPANA analyses the collected data by using the programs which read the
spana_lsq.dta file. For this purpose, the general least square analyses
programs written in C language, spana_lsq.c, is provided in the version
5.
The user prepares C-source file in which the function describing the user's
system is defined, links it with spana_lsq.c, and compiles to make the
execute program of the ".exe" type, which may be called from
SPANA directly. For compilation of the programs, not only commercial C
compilers but also free one such as DJGPP and Micosoft Visual Studio C++ are available.
The following variables, parameters, and constants are defined ;
y[i] : dependent variables (ex. Abs in the titration experiment), i =
1 - 20
x[i] : independent variable (ex. titrant concentration), i = 1 (fixed in SPANA)
c[i] : constants (ex. sample concentration), i = 1 - 30
p[i] : optimized parameters (ex. equilibrium constants, molar absorption
constants), i = 1 - 20
By using these symbols, the functions of user's system, y[i] = f (x[i]
, c[i] , p[i] ), are defined in the C-source as
void function( int i, double x[], double y[])
{ }
(where i is number of dependent variables).
The following program is that applicable for determination of thermodynamic
parameters, ΔH and ΔS. by using temperature dependent spectral titration
data of the 1:1 complex formation system at 10 different wave length position.

In this source program, the lines between /* and */ are comments and the
following part defines the difference absorption,
ΔAbs = (Abs - εA[A] 0 =) Δε[AB].
After saving this file, for example, as the name of "titrn_t-dep.c",
the execution program, titrn_t-dep.exe, is obtained by linking with "spana_lsq.c"
and compiling the source programs as follows,
gcc -o titrn_t-dep.exe spana_lsq.c titrn_t-dep.c, for DJGPP
or
cl titrn_t-dep.c spana_lsq.c, for Visual Studio C++
SPANA ver.5 provide another least square program for the analysis of the
reaction kinetics, spana_redap.c, which analyses the time dependent spectral
change by numerical integration of differential equations for chemical
reactions without using the integral forms. For the detail of this program,
contact me by e-mail.